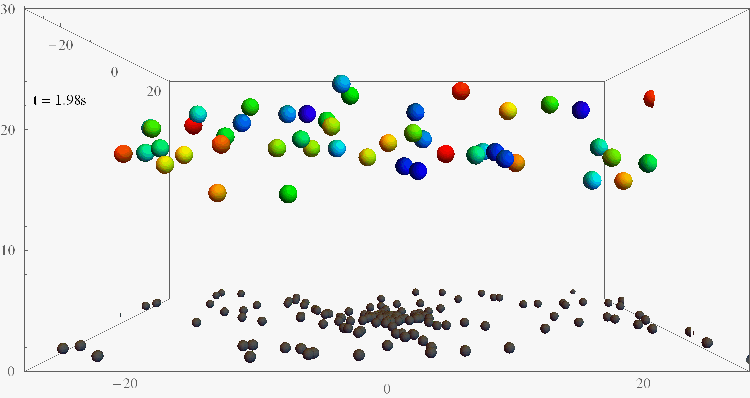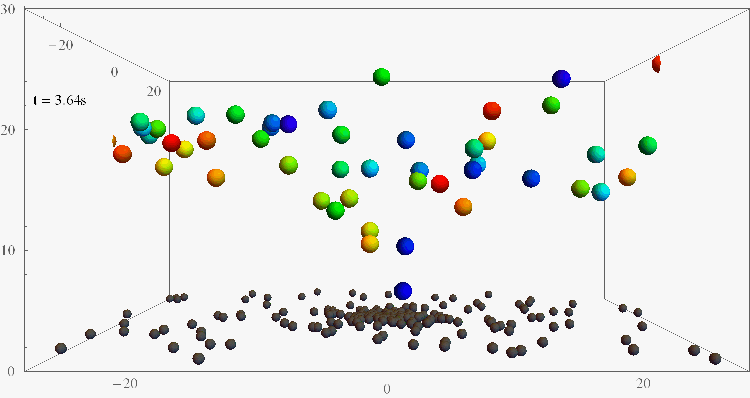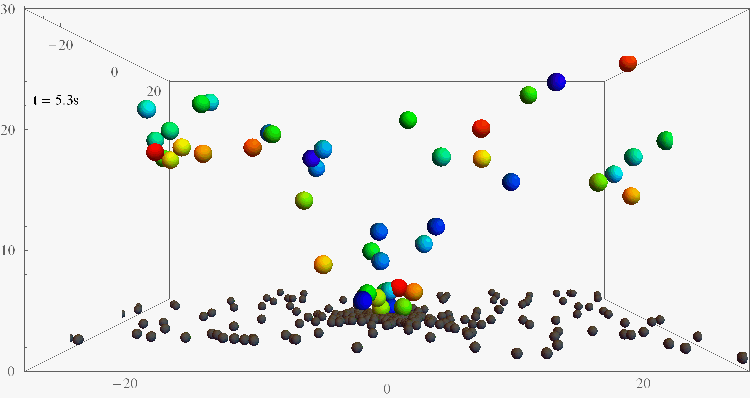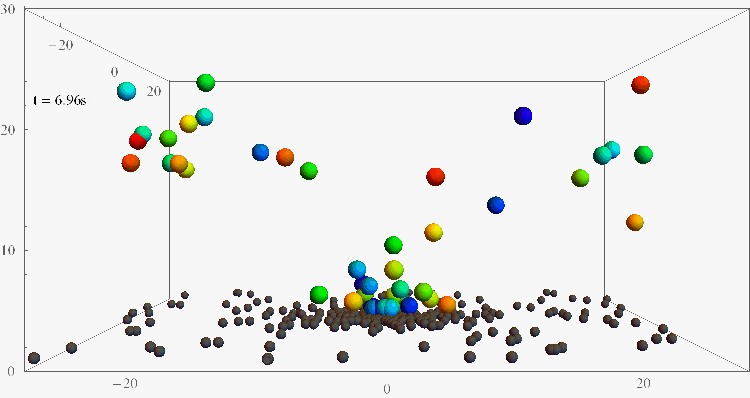Research
Physics of Pathogens
As a more applied branch of research, we study the physics of pathogen dynamics. Indeed, besides the complex repertoire of biochemical interactions, microbial dynamics strongly depend on biophysical factors. Especially fluid mechanics are important, because both biological micro-organisms and synthetic micro-robots must propel through viscous liquids to achieve their goal, be it to invade new territories or to deliver drugs to infected regions. Considerable attention is devoted to learning how to prevent or encourage these processes, and understanding the interactions between micro-swimmers and their complex environments is an essential part of this. In vivo conditions provide a challenge to model, although novel experimental, computational and theoretical techniques have provided clear insights into the continuous interplay between the effects of strong confinement, hydrodynamic interactions, and local activity that drives living systems out of equilibrium.
Mucociliary clearance in the respiratory system
Flows generated by cilia in the mouse airway (Nature Physics, 2020)
Upstream swimming of bacteria
Bacterial contamination of biological channels, catheters or water resources is a major threat to public health, which can be amplified by the ability of bacteria to swim upstream. The mechanisms of this “rheotaxis”, the reorientation with respect to flow gradients, are still poorly understood. To understand these dynamics, we follow individual E. coli bacteria swimming at surfaces under shear flow with two complementary experimental assays, based on 3D Lagrangian tracking and fluorescent flagellar labelling, and we develop a theoretical model for their rheotactic motion. This research could shed new light on bacterial transport and reveal new strategies for contamination prevention, rheotactic cell sorting, and microswimmer navigation in complex flow environments.
Bacteria swimming against the flow
(Nature Comm 2019, PNAS 2024)
Intelligent Active Matter
Since Erwin Schrödinger published “What is life?“, the puzzle of understanding and predicting how biological systems emerge from the collective dynamics of their constituents remains standing. This puzzle spans across the scales: The combined efforts of molecular motors give rise to the dynamics of a cell, the shared functionality of cells leads to unicellular colonies or mammalian tissues that together form organs and organisms, which in turn demonstrate cooperative behaviour in self-assembled flocks, swarms and schools. The field of “active matter” has developed into a major interdisciplinary science community to comprehend this upward cascade, but the potential of active materials to sense, communicate and compute information remains widely unexplored.
Indeed, a fundamental aspect of life is the ability to process information. On the one hand, responding quickly to stimuli is key to survival, which begs the question how fast can organisms react within the laws of physics? On the other hand, information feedback is at the heart of any autonomous and self-replicating machine, so the profound relationship between sensors and actuators has also been essential to the development of complex life forms. But how ‘complex’ must a physical system be in order to show life-like phenotypes that resemble information processing, memory storage, or “intelligence”? Here we aim to approach these questions by combining concepts from cell biology, physics and information theory.
Hydrodynamic communication between cells (Nature, 2019)
Hydrodynamic trigger waves
The biophysical relationships between sensors and actuators have been fundamental to the development of complex life forms. We recently discovered a form of rapid hydrodynamic communication in the unicellular protist Spirostomum ambiguum. By contracting its long body within milliseconds, these cells can accelerate up to 14g-forces to produce long-ranged flows. Neighbouring organisms can sense these flows, which leads to hydrodynamic trigger waves that quickly travel from one cell to the next, at speeds a thousands times faster than the cell motility. Hence, small signals are amplified across entire colonies. We are interested in the biological functions of such mechanical signals, across different organisms, and how cells can survive this ultra-fast motion in the first place.
Motile active matter
Bacteria aggregate on surfaces to form dense communities that can develop into biofilms. This crowding drains their food reserves rapidly but, surprisingly, we discovered that ‘active carpets’ of motile microbes can replenish resources by creating strong currents that transport nutrients and oxygen. By swimming each bacterium produces a small flow, but together these whorls are amplified when the cells organise themselves into specific patterns. Large vortices then emerge, which can span distances even beyond the colony size. We calculate the flow strengths generated by these patterns and predict that this co-operative nutrient transport can sustain microbial activity. Understanding how single cells assemble to function collectively is a fundamental pillar of biology. Conversely, new technology to disturb these replenishing flows is immediately applicable to preventing bacterial biofilm formation, reducing the need to apply biocides and lowering chances of infection.
Micro-robotics
As highlighted by the Nobel laureate Edward Purcell in his famous talk “Life at low Reynolds number“, the motion of fluids at microscopic length scales can be very counter-intuitive. Where a physicist would write that a Force is proportional to Accelleration, F∝a (Newton’s second law), a microbiologist would write that a Force is proportional to Velocity, F∝v (Stokes’ law). This difference has important consequences: For example, time does not matter for flows at low Reynolds numbers, only configurations. Therefore, a cell that stops swimming will come to a halt effectively instantaneously, indeed because the viscous forces are much larger than inertia. Similarly, it is very difficult to mix microfluids, because if you stir in the opposite direction everything ends up where it started earlier. While powerful mathematical tools have been developed to solve micro-flows, there is still a wealth of problems to tackle, especially in combination with ideas from physics and biology.
Active materials & particle delivery
Mobile microrobots are expected to transform bioengineering and therapeutics by precisely navigating in microchannels and performing minimally invasive treatments. The functionalities of these magnetic microrobots have grown substantially in recent years. However, delivery at specific target locations remains challenging. Freely swimming microrobots have relatively low speeds, and it is difficult to swim against the complex fluid flows inside the human body. We aim to develop a new microscale biomedical delivery mechanisms that overcomes these challenges.
Since the Hollywood movie “Fantastic Voyage”, mobile micromachines that can navigate inside the body and cure diseases have been a Holy Grail for scientists and engineers. Despite the advances in imaging, materials, actuation, control, and navigation, it is still very challenging due to the high complexity of in vivo environments and low reliability of these processes. By introducing artificial microtubules, we believe these “micro-highways for microrobots” can provide an alternative solution to the free-swimming microrobots, bringing robust biomedical microtransport much closer to reality.
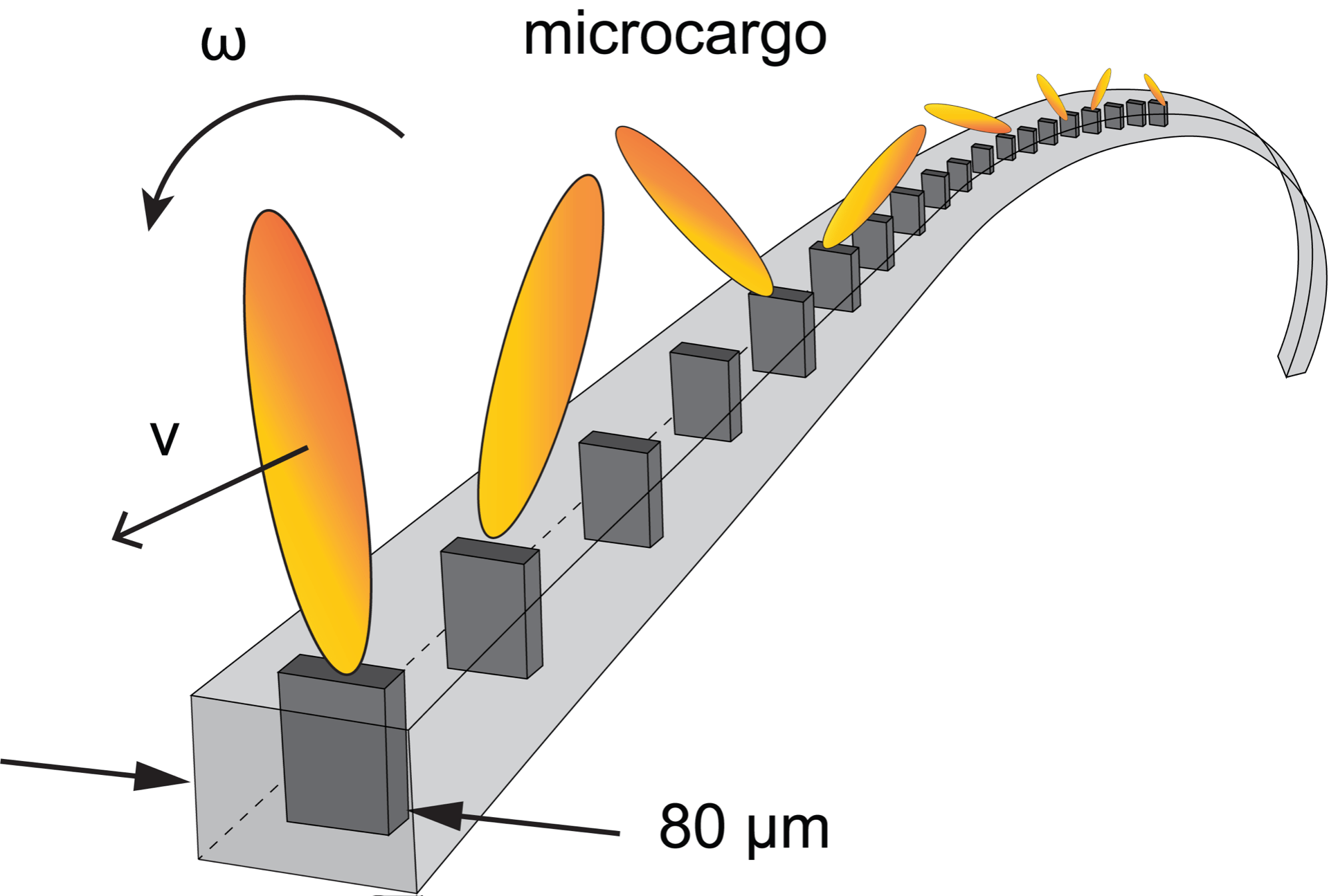
Artificial cilia & self-cleaning surfaces
One of the major challenges in modern robotics is controlling micromanipulation by active and adaptive materials. In the respiratory system, such actuation enables pathogen clearance by means of motile cilia. While various types of artificial cilia have been engineered recently, they often involve complex manufacturing protocols and focus on transporting liquids only. Here, soft magnetic carpets are created via an easy self-assembly route based on the Rosensweig instability. These carpets can transport not only liquids but also solid objects that are larger and heavier than the artificial cilia, using a crowd-surfing effect.This amphibious transportation is locally and reconfigurably tunable by simple micromagnets or advanced programmable magnetic fields with a high degree of spatial resolution. Two surprising cargo reversal effects are identified and modeled due to collective ciliary motion and nontrivial elastohydrodynamics. While the active carpets are generally applicable to integrated control systems for transport, mixing, and sorting, these effects can also be exploited for microfluidic viscosimetry and elastometry.
Swimming microrobots
Because the Reynolds number is very small for microswimmers, they must use special locomotion strategies in order to move forwards at all. Moreover, because of the energy that swimmers produce continually at the smallest length scales, their motion cannot be described by classical thermodynamics since the system is driven inherently out of equilibrium. As a result, many unexpected phenomena can occur in swimmer systems, including giant number fluctuations, rectification, and collective motion.
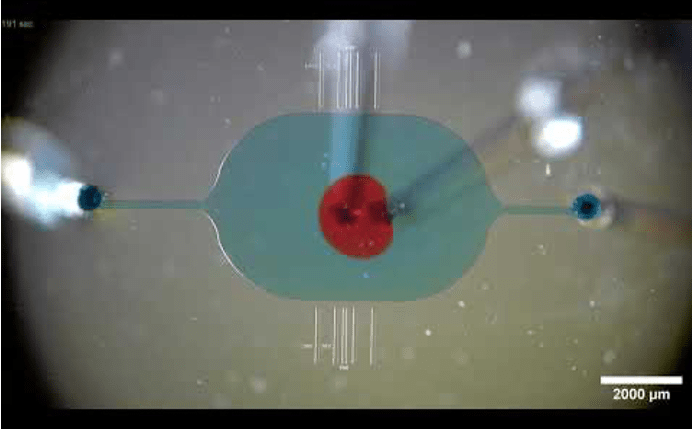

A microfluidic channel for particle sorting
Microfluidics and sorting microplastics
The ability to separate microscopic objects lies at the core of many important applications. First, sorting microplastics is crucial for preventing pollution of our marine ecosystems, which gets harder and harder as they degrade to smaller particles. Second, separating cells is also essential in biomedical research, for diagnosing diseases like malaria or chronic fatigue syndrome, and for DNA sequencing of individual cells. Modern lab-on-a-chip instruments (also known as microfluidic devices) now provide an avenue for low-cost sorting, which can facilitate early-stage diagnosis and improve treatments worldwide. However, a persistent problem is clogging, as biomaterials stick to the internal chip surfaces, which makes these tools inefficient or unreliable. We aim to introduce new techniques that could solve these clogging issues, and to develop new instruments that can be of use in many branches of biomedicine. We are also developing a program for outreach and interdisciplinary science education for in-classroom demonstrations of sorting microplastics.